By Qing Xu //
G lioblastoma multiforme (GBM) is one of severe brain tumours. It is characterized by high mortality and median survival time of patients is one year. Manual analysis of histopathology images is regarded as a traditional method to identify disease stage. However, it has some shortages such as different evolution criteria of experts and long wait for result. In this project, an approach combined with deep learning and image processing has been proposed. It shows powerful ability to detect and classify different cells as well as provides a result of proliferation index to aid clinician’s diagnosis.
lioblastoma multiforme (GBM) is one of severe brain tumours. It is characterized by high mortality and median survival time of patients is one year. Manual analysis of histopathology images is regarded as a traditional method to identify disease stage. However, it has some shortages such as different evolution criteria of experts and long wait for result. In this project, an approach combined with deep learning and image processing has been proposed. It shows powerful ability to detect and classify different cells as well as provides a result of proliferation index to aid clinician’s diagnosis.
There are some challenges we faced during this project. Firstly, some of the popular neural networks, such as Faster-RNN and FCN, could not achieve expected performance due to the small datasets containing only 22 original images. In order to extend datasets, we decided to extract various cells from each image and used multiple image augmentation. In this way, we got considerable samples and successfully trained an efficient network using ResNet50 at the end.
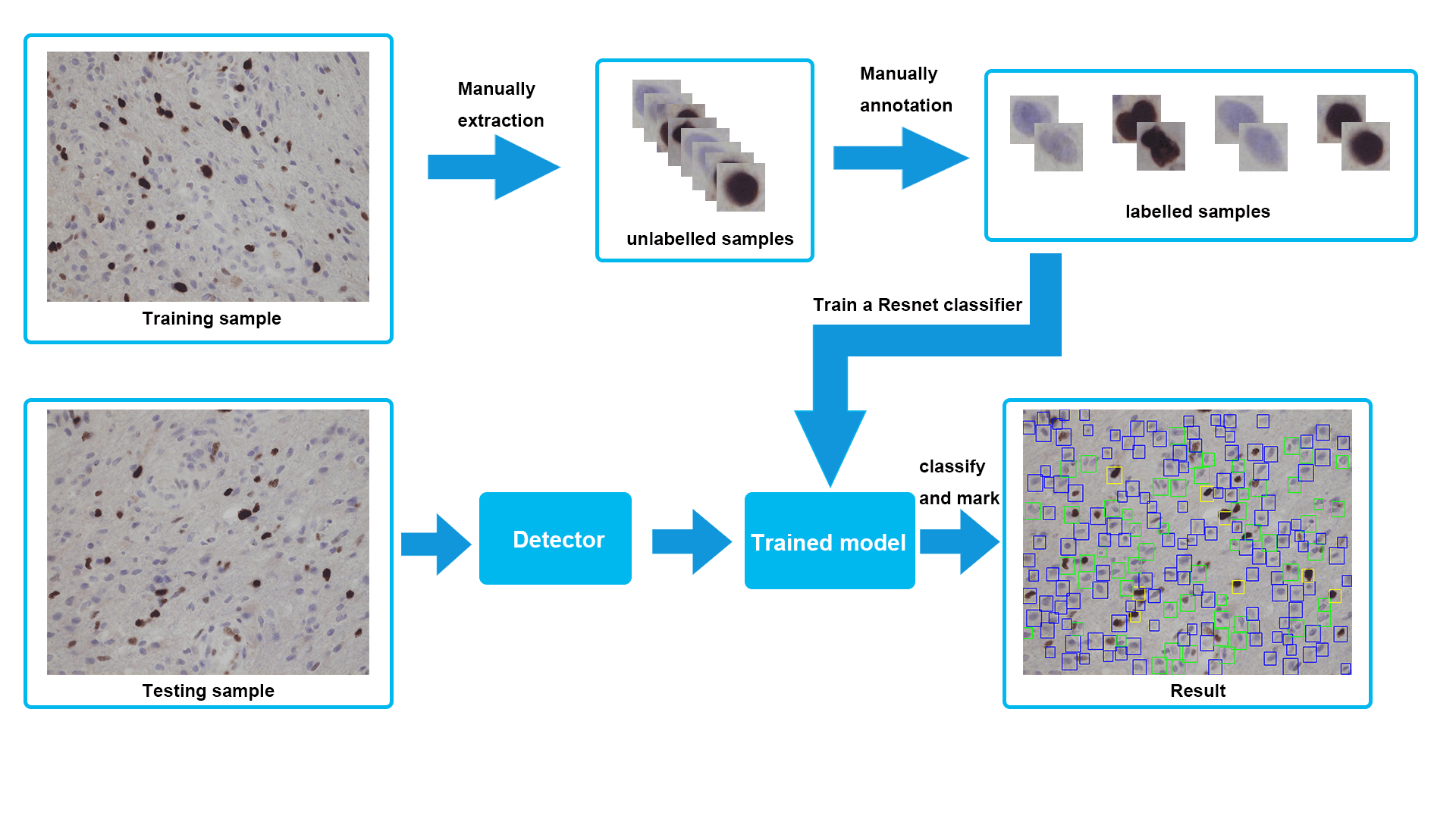 Another obstacle was that our designed detector showed a low accuracy in terms of recognising different cells at the beginning. Specifically, with the background often identified as cells. When we checked datasets, we found only four different cell categories were included, without background, and some of training and validating samples were misclassified. Therefore, we started to collect some samples of the background, fixed incorrect datasets and trained a new model. The final model demonstrated a high performance on the task of classification and detection. One of testing result has been presented in Figure 1.
Another obstacle was that our designed detector showed a low accuracy in terms of recognising different cells at the beginning. Specifically, with the background often identified as cells. When we checked datasets, we found only four different cell categories were included, without background, and some of training and validating samples were misclassified. Therefore, we started to collect some samples of the background, fixed incorrect datasets and trained a new model. The final model demonstrated a high performance on the task of classification and detection. One of testing result has been presented in Figure 1.
The result of this particular challenge aided my understanding of how diverse datasets can affect the result of classification. If we prepare wrong datasets, the trained model will be hard to achieve our aim in testing stage, and although it indicates a high training and validating accuracy during the training stage. Throughout this project, I have learned how to gather demanded data, construct a network framework using Python and PyTorch; and evaluate performance of a trained model.
I had two supervisors who supported me during this project. Both Dr Wenting Duan and Prof Xujiong Ye were happy to answer any question and concerns I had and welcomed my input to the project. Although it was difficult to meet up face-to-face due to the outbreak of the Covid-19, both supervisors were able to provide regular constructive progress updates by email or other methods. I cannot thank Wenting and Xujiong enough for this experience.
UROS has provided me with the opportunity to gain a high-level research experience and further aiding my understanding of the whole research process. This research has helped to direct me to the area which I am interested in and want to study in the future. Based on the experience of this project, I have decided to continue the research of medical images processing that combined with deep learning in my further Masters degree.
*To view Qing’s research poster and presentation recording, please click on the thumbnails below: 
